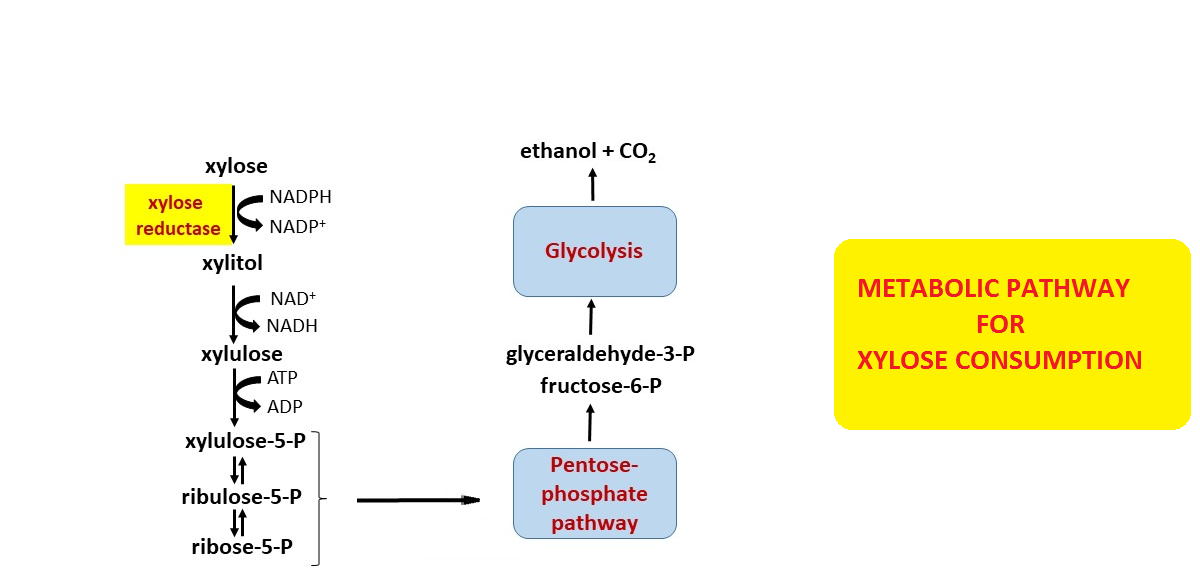
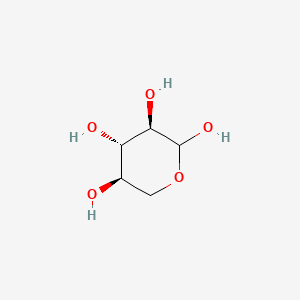
The metabolic pathway for production of ethylene glycol from D-xylose.... | Download Scientific Diagram

UNIVERSITÀ DEGLI STUDI DELLA TUSCIA DI VITERBO DIPARTIMENTO PER L INNOVAZIONE NEI SISTEMI BIOLOGICI AGROALIMENTARI E FORESTALI - PDF Free Download
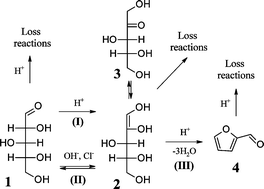
Chloride ions enhance furfural formation from d-xylose in dilute aqueous acidic solutions - Green Chemistry (RSC Publishing)

d‐Xylose consumption by nonrecombinant Saccharomyces cerevisiae: A review - Patiño - 2019 - Yeast - Wiley Online Library
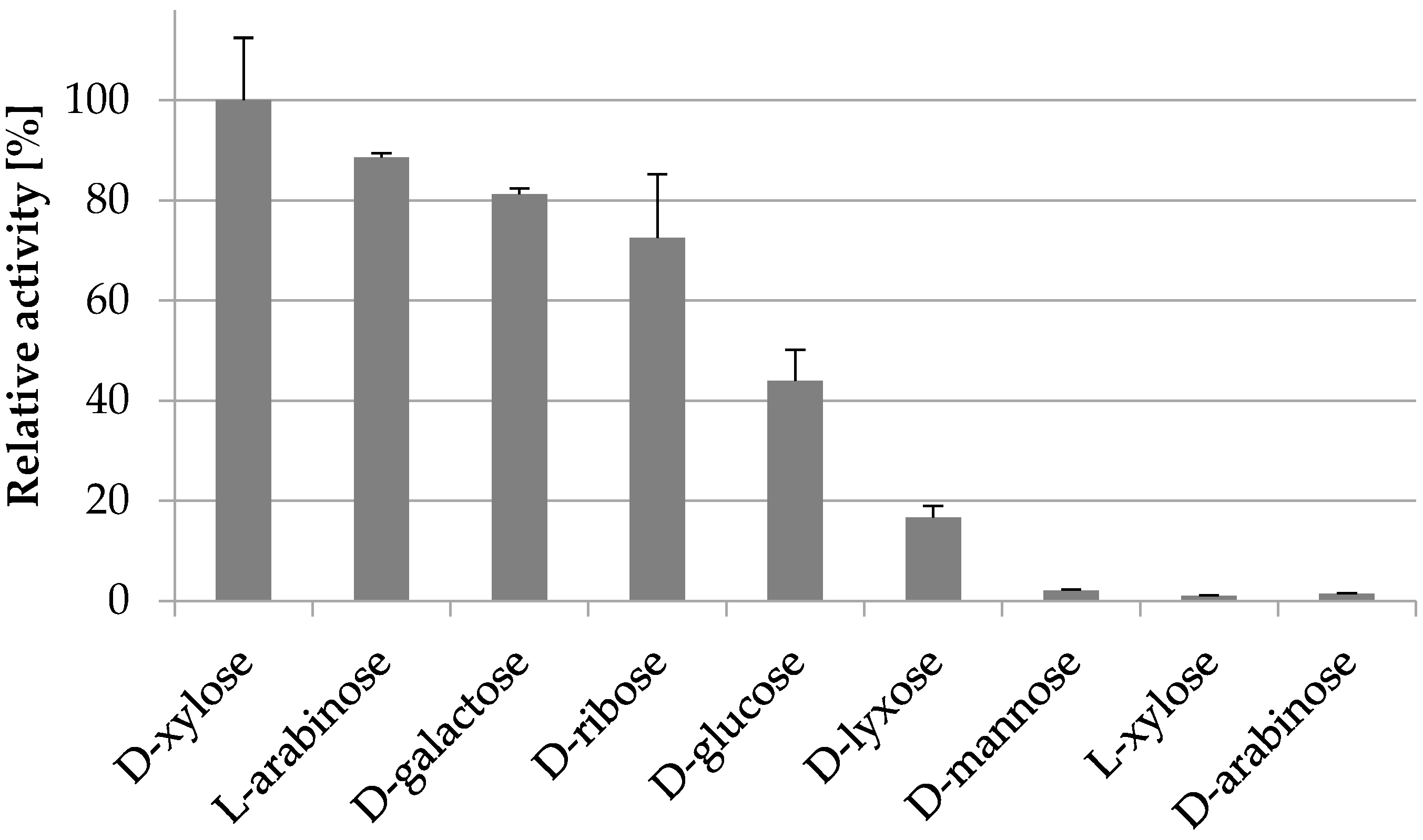
IJMS | Free Full-Text | Kinetics and Predicted Structure of a Novel Xylose Reductase from Chaetomium thermophilum | HTML

A Novel Aldo-Keto Reductase, HdRed, from the Pacific Abalone Haliotis discus hannai, Which Reduces Alginate-derived 4-Deoxy-l-erythro-5-hexoseulose Uronic Acid to 2-Keto-3-deoxy-d-gluconate* - Journal of Biological Chemistry

Synthetic (blue) and natural (black) d-xylose assimilation pathways. In... | Download Scientific Diagram

L-Arabinose Binding, Isomerization, and Epimerization by D-Xylose Isomerase: X-Ray/Neutron Crystallographic and Molecular Simulation Study: Structure
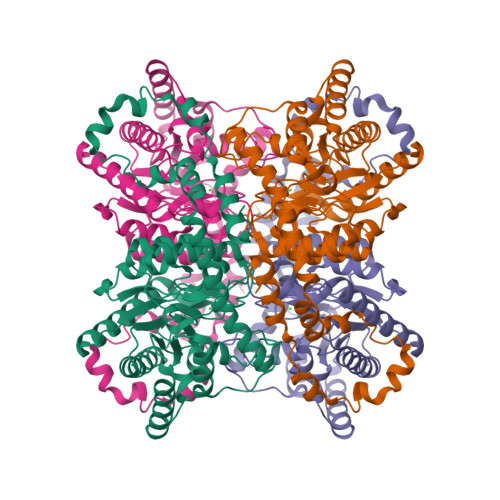
RCSB PDB - 3KBN: Room temperature structure of D-Xylose Isomerase in complex with 2Ni(2+) co-factors and d12-D-glucose in the linear form

L-Arabinose Binding, Isomerization, and Epimerization by D-Xylose Isomerase: X-Ray/Neutron Crystallographic and Molecular Simulation Study: Structure